

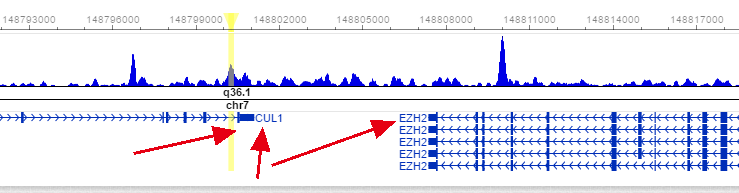
We observed spatio-temporal activity of TLR9 and TLR3 specific enhancers regulating signal specific target genes. We profiled gene expression changes along with H3K27ac active enhancers and NCoR1 binding in the TLR9, TLR3 and combined TLR9 + TLR3 activated cDC1.

In this study, we did a comprehensive epigenomic analysis of murine conventional type-I DCs (cDC1) across different TLR ligation conditions. However, how NCoR1 regulates enhancer activity and gene expression in individual or multiple Toll-like receptor (TLR) activation in DCs is largely unknown. Apart from transcription factor binding, dynamic regulation of enhancer activity through global transcriptional repressors like Nuclear Receptor Co-repressor 1 (NCoR1) plays a major role in fine-tuning of DC responses. It will only tell you where the regions that you have identified are located even though there is more intergenic regions in the genome.Tight control of gene regulation in dendritic cells (DCs) is important to mount pathogen specific immune responses. Now keep in mind that what you probably want to know is enrichment over genomic background and this figure will not tell you that. It is also possible to make higher quality figures by copying to pdf and in dev.copy() and converting the pdf into png using the convert program on linux. The figures thus far look small and cramped, you can export the figures are a bigger size usingĭev.copy(png, "mypeaks.png", width=800, height=600) dev.off() Īfter some cropping, this is the same picture as the one shown in the beginning of the post. Pie(newtable, main="mypeaks annotation", col=rainbow(9)) Next, I will choose better looking colors than the default, this can be done in two ways, either specifying the colors or choosing colors from gradient, I use the latter: Next, I will add %s and numbers to each of the labels: Pie( newtable, main="mypeaks annotation") To solve this, we can initalize a table of 0s called newtable. Now if you look closely, you will notice that this pie chart is missing a category, on the HOMER annotation page, it lists 7 categories but the above chart is only showing 6 (missing promoter-TSS).
Pietable = table(unlist(lapply(strsplit(as.character(anno$Annotation), " \\("),"[[",1))) #take the part before first ' (' By cutting this out, we can create a pie chart. You will notice that the 8th row (Annotation) conetains information on where the peak overlaps. Wingless-type MMTV integration site family, member 10AĪnno = read.table("anno_mypeaks.txt", sep="\t", header=T, quote="") This will give you a file that looks like this: I ran something similar:Ī mypeaks.bed hg19 > anno_mypeaks.txt However, pie charts are still often used in biology to display this kind of information.įirst you will want to generate annotation data outlined on this page of HOMER documentation. A bar chart or dot chart is a preferable way of displaying this type of data. The eye is good at judging linear measures and bad at judging relative areas. Pie charts are a very bad way of displaying information. This post will go through the steps create a pie chart of genomic annotations output by HOMER using Rįirst, I would like to begin with a disclaimer from the documentation of the pie() function in R: HOMER is a set of programs that can be used to call and annotate chip-seq peaks (among other things). HOMER is typically used to analyze chip-seq/faire-seq data. This post will go through the steps to make a pie graph annotating binding locations using HOMER output.
